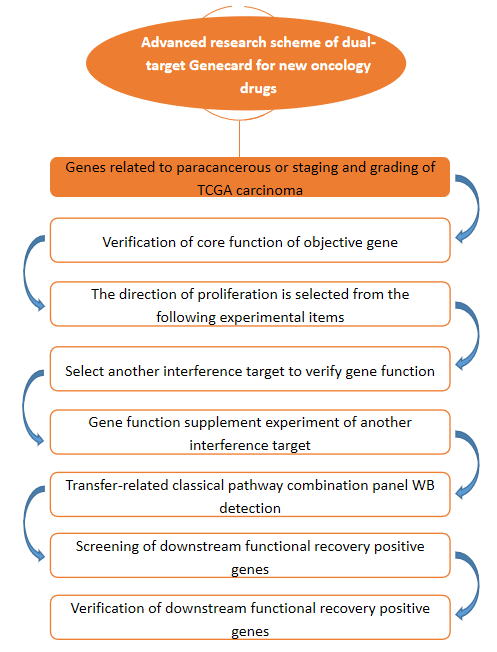
Dual-Target Research Scheme of New Tumor Drug Target Genecard
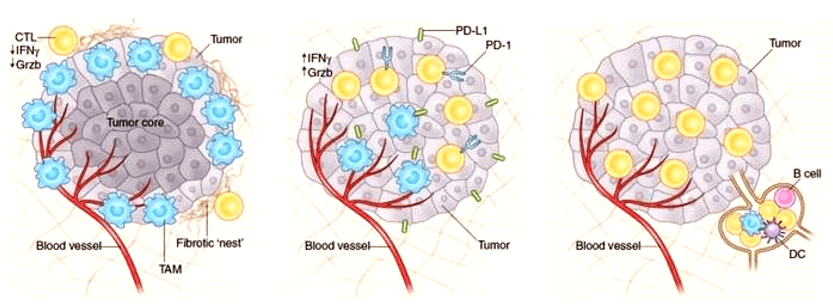
Through TCGA big data, we analyzed the high expression or expression of genes related to disease staging and grading in cancer tissues, and analyzed the candidate genes through the disease key gene database we established. Using lentivirus-mediated RNAi technology, several candidate genes were RNAi, through Cellomics high connotation function screening (HCS) platform, and the infected cells were counted for 4 to 5 days. The genes affecting cell proliferation were found. The positive results of at least three proliferation functional experiments (MTT/ cell cycle (PI/ flow) / apoptosis (Annexin V / flow) / BrdU/ clone formation, etc.) and two metastatic functional experiments (scratch healing / Transwell migration / Transwell invasion, etc.) of the two interference targets of the gene on one cell line were obtained. After interfering with the target gene in a cell line, the transfer-related classical pathway combination panel* was verified by WB. According to the results of WB panel, a positive mechanism gene was selected to construct a virus that interfered with or overexpressed the target gene on the cell line, and interfered with or overexpressed the downstream mechanism gene on the cell line. The changes of cell proliferation ability were observed by HCS scanning for 5 days to verify the functional recovery effect of the downstream mechanism gene on the target gene. The positive response results of a proliferation-related and a metastasis-related functional experiment were obtained.
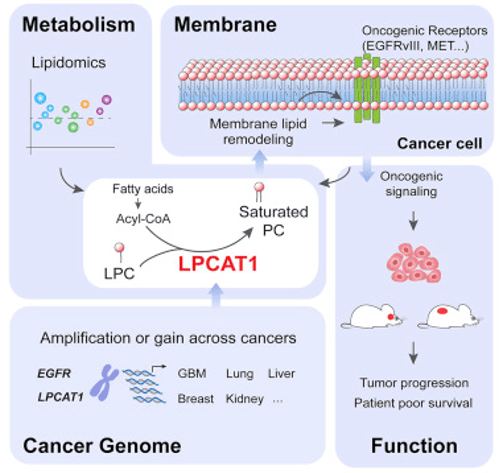 Figure 1. Amplification of oncogenes in the growth factor signaling pathway leads to cancer dependent on membrane lipid remodeling(Junfeng Bi, et al. 2019)
Figure 1. Amplification of oncogenes in the growth factor signaling pathway leads to cancer dependent on membrane lipid remodeling(Junfeng Bi, et al. 2019)
Transfer-related Classical Pathway Combinations Panel
AKT1, p-AKT (Ser473 or Thr308), mTOR, p-mTOR (Ser2448), ERK1/2, p-ERK1/2 (Thr202+Tyr204), P38, p-P38 (T180+Y182), MYC, β-Catenin, p-β-Catenin (S33+S37), NFkB-p65, p-NFkB-p65 (Ser536), CDH1 (E-Cadherin), CDH2 (N-Cadherin), MMP2, MMP9, FN1, VIM, Snail, Slug, TWIST and other 22 genes.
Service Flow
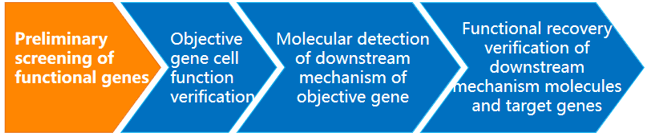

Our Advantage
Mature and reliable experimental analysis platform of Genecard dual-target research scheme: rich project experience, participation in major projects, high-level cooperation results published.
Based on high-quality data analysis, pre-research and verification of the contents and methods of analysis, it is indeed mature and feasible.
Creative Proteomics leads a rich team that uses the most rigorous experimental solutions and internationally recognized experimental techniques for Genecard dual-target research scheme to obtain your customized information. Use the most accurate experimental results to help you better carry out tumor-related scientific research.
Reference
1. Junfeng Bi, et al. (2019). "Oncogene Amplification in Growth Factor Signaling Pathways Renders Cancers Dependent on Membrane Lipid Remodeling." Cell Metabolism.
* For Research Use Only. Not for use in the treatment or diagnosis of disease.
Related Services:


 Figure 1. Amplification of oncogenes in the growth factor signaling pathway leads to cancer dependent on membrane lipid remodeling(Junfeng Bi, et al. 2019)
Figure 1. Amplification of oncogenes in the growth factor signaling pathway leads to cancer dependent on membrane lipid remodeling(Junfeng Bi, et al. 2019)