There are practical or potentially valuable cancer single cell research methods in cancer research, including single cell isolation, whole genome and transcriptome amplification, epigenome analysis, multidimensional sequencing and next generation sequencing and analysis. Cancer single cell isolation is the most basic single cell experiment. The analysis of individual cancer cells initially requires the isolation of the desired type of cells based on classical phenotypes or surface markers in a manner that maintains the biological integrity of the cells. When isolating a single cell from a solid tumor, the tumor tissue needs to be dispersed into a single cell suspension; there are two methods involved in isolating carbon tetrachloride from body fluids; and the CTC capture platform CTC-IchIP, which depends on the inherent difference in shape or size between white blood cells and CTC, can be used to isolate CTC from which no known biomarkers are known. For DTC isolation, needle aspiration was used to collect bone marrow and then single cell sorting was performed. Creative Proteomics leads the best team to bring you better experimental results of tumor single cell isolation and culture, and help you better study tumor.
Experimental Method
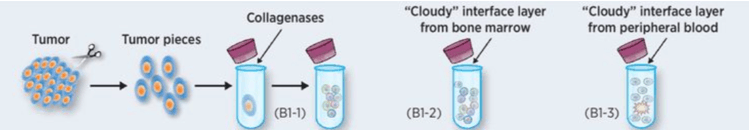 Figure 1. Preparation of tumor single cell suspension (Zhang X, et al. 2016)
Figure 1. Preparation of tumor single cell suspension (Zhang X, et al. 2016)
First, the tumor tissue was placed on a microslide and the surface was covered with a thermoplastic film. The laser passes through the film and focuses on the target cell, which is attached to the cell and separates the cell from the tissue by mechanical displacement. Next, when the single cell is isolated from the solid tumor, the tumor tissue is dispersed into the single cell suspension. Mechanical separation and enzyme treatment are often used (figure 1).
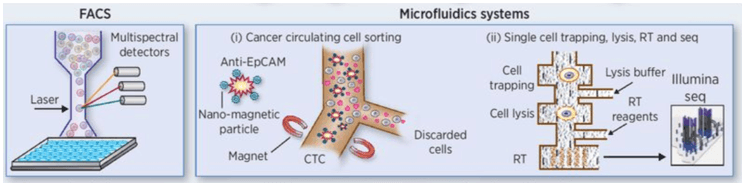 Figure 2. Common method of tumor single cell isolation: FACS, microfluidic system (Zhang X, et al. 2016)
Figure 2. Common method of tumor single cell isolation: FACS, microfluidic system (Zhang X, et al. 2016)
The separation of carbon tetrachloride from body fluids involves two methods (figure 2, 3). The first is biomarker-dependent, such as the CellSearch magnetic bead system. The second uses density gradient centrifugation, microfiltration or microfluidic systems. CTC capture platform CTC-IchIP, which depends on the inherent difference in morphology or size between leukocytes and CTC, can be used to isolate some CTC with no known biomarkers. For DTC isolation, needle aspiration was used to collect bone marrow and then single cell sorting was performed.
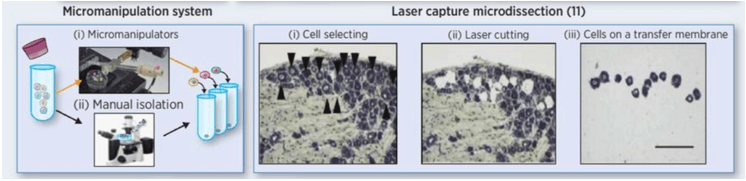 Figure 3. Common methods of tumor single cell isolation: micromanipulation system and LCM (Zhang X, et al. 2016)
Figure 3. Common methods of tumor single cell isolation: micromanipulation system and LCM (Zhang X, et al. 2016)
Single cell sorting methods, including marker / phenotypic methods for isolating single cells from large cell populations [FACS, microfluidic, micromanipulation, and LCM and rare cell populations (figure 2, 3). Tissues or organs (such as intact tumors) unbiased, unlabeled decomposition into different types of single cells can omit the FACS process, but still need to dissociate the tissue into single cells or locate single cells, usually need to analyze a large number of cells.
Our Advantage
Creative Proteomics's single-cell separation technology can produce 100, 000 single-cell libraries in a single experiment, and users can customize primers according to their needs to focus on the target genes, thus greatly reducing the cost of subsequent sequencing.Suitable for tissues, cell lines, can enrich target cells through specific antibodies.
Creative Proteomics leads a rich team that uses the most stringent data quality control standards and internationally recognized experimental techniques to analyze tumor single cell isolation methods for your customized information. to help you effectively screen and systematically analyze the heterogeneity of these different cancer cells and cancer cells.
Reference
1. Zhang X, Marjani S L, Hu Z, et al. Single-cell sequencing for precise cancer research: progress and prospects. Cancer Research, 2016, 76(6): 1305-1312.
* For Research Use Only. Not for use in the treatment or diagnosis of disease.
Related Services:

 Figure 1. Preparation of tumor single cell suspension (Zhang X, et al. 2016)
Figure 1. Preparation of tumor single cell suspension (Zhang X, et al. 2016) Figure 2. Common method of tumor single cell isolation: FACS, microfluidic system (Zhang X, et al. 2016)
Figure 2. Common method of tumor single cell isolation: FACS, microfluidic system (Zhang X, et al. 2016) Figure 3. Common methods of tumor single cell isolation: micromanipulation system and LCM (Zhang X, et al. 2016)
Figure 3. Common methods of tumor single cell isolation: micromanipulation system and LCM (Zhang X, et al. 2016)