Cancer research aims to focus on understanding the causes, progression, and deteriorating effects of cancer on the patient in the ultimate goal to develop safe and effective methods to diagnose, prevent, and cure these diseases. Researches in the past have made a great impact on early detection, screening, and developing treatments for cancer patients. One of the helpful technologies that researchers use is metatranscriptomics. Metatranscriptomics is a branch of molecular research that focuses on the gene expression of biological entities, such as bacteria and viruses, found in a particular environmental sample. It gives information on the function and activity of genes in the microbial community and its interaction with other organisms such as humans- normal microflora interactions. Compared to metagenomics, which provides an idea on the microbial community present in the sample, metatranscriptomics exposes the function and activity of transcripts, profiles the expression of these genes, and shows reactions and interactions of these microorganisms and their host to show their effects on the gene expression of theirs. Studies have also been conducted in the past using metatranscriptomic sequencing in order to determine pathogen and host interaction and its impact on the development of certain diseases, including cancer.
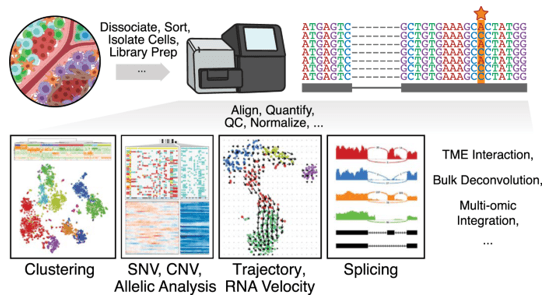 Figure 1. Single cell transcriptomics sequencing workflow and downstream computational analyses. (Fan 2020)
Figure 1. Single cell transcriptomics sequencing workflow and downstream computational analyses. (Fan 2020)
Studies have used metatranscriptomics in conjunction with metagenomics to map out the microorganisms present in clinical samples and to analyze their activity in order to determine which causes damage to cells or which induces the development of tumors. This technique also provides information on host cell expression as a response to the infection. Feng, et al. (2019), used metagenomic and metatranscriptomic analysis in determining human microflora and host cell interaction in the development of prostate cancer. The information they gathered using this technology provided potential preliminary evidence regarding the influence of normal microflora to the development of diseases and tumors. They found that the expression levels of certain microbial genes influence the expression of host genes leading to the abnormal production of resulting proteins that could be linked to the progression of cancer in the prostate. Metatranscriptomics sequencing also provides information on potential microbial therapeutic targets which can help in designing effective treatments for a specific type of cancer considering also the specificity of treatment per individual patient.

Metatranscriptomics has proven its importance in cancer research to understand the behavior of the host. Creative Proteomics offers metatranscriptomic sequencing services, from sample preparation up to bioinformatics analysis, to aid in your cancer research. Following the basic workflow above, samples shall undergo RNA isolation, purification, and quantification as part of sample preparation. Then RNA libraries will then be prepared before sequencing. Sequencing can either be done using Illumina or PacBio sequencing technologies, after which profiling and analysis will be done.
Creative Proteomics is one of the most trusted companies in metatranscriptomics and cancer research, we are dedicated to providing the best strategies for your researches. With state-of-the-art instrumentations, we provide fast and reliable metatranscriptomics sequencing services. Let us help you make a change in the world. For additional information and other services that we provide, please feel free to contact us.
References:
1. Fan, J., Slowikowski, K. Zhang, F. Single-cell transcriptomics in cancer: computational challenges and opportunities. Exp Mol Med. 2020.
2. Feng, Y., Ramnarine, V.R., Bell, R., et al. Metagenomic and metatranscriptomic analysis of human prostate microbiota from patients with prostate cancer. BMC Genomics. 2019, 20:146.
3. Peters, B.A., Wilson, M., Moran, U., et al. Relating the gut metagenome and metatranscriptome to immunotheraphy responses in melanoma patients. Genome Medicine. 2019, 11:61.
4. Shakya, M., Lo, C., Chain, P.S.G. Advances and Challenges in Metatranscriptomic Analysis. Frontiers in Genetics. 2019, 10:904.
5. Warnecke F., Hess M. A perspective: metatranscriptomics as a tool for the discovery of novel biocatalysts. Journal of Biotechnology. 2009, 142(1): 91-95.
* For Research Use Only. Not for use in the treatment or diagnosis of disease.
Related Services:

 Figure 1. Single cell transcriptomics sequencing workflow and downstream computational analyses. (Fan 2020)
Figure 1. Single cell transcriptomics sequencing workflow and downstream computational analyses. (Fan 2020)